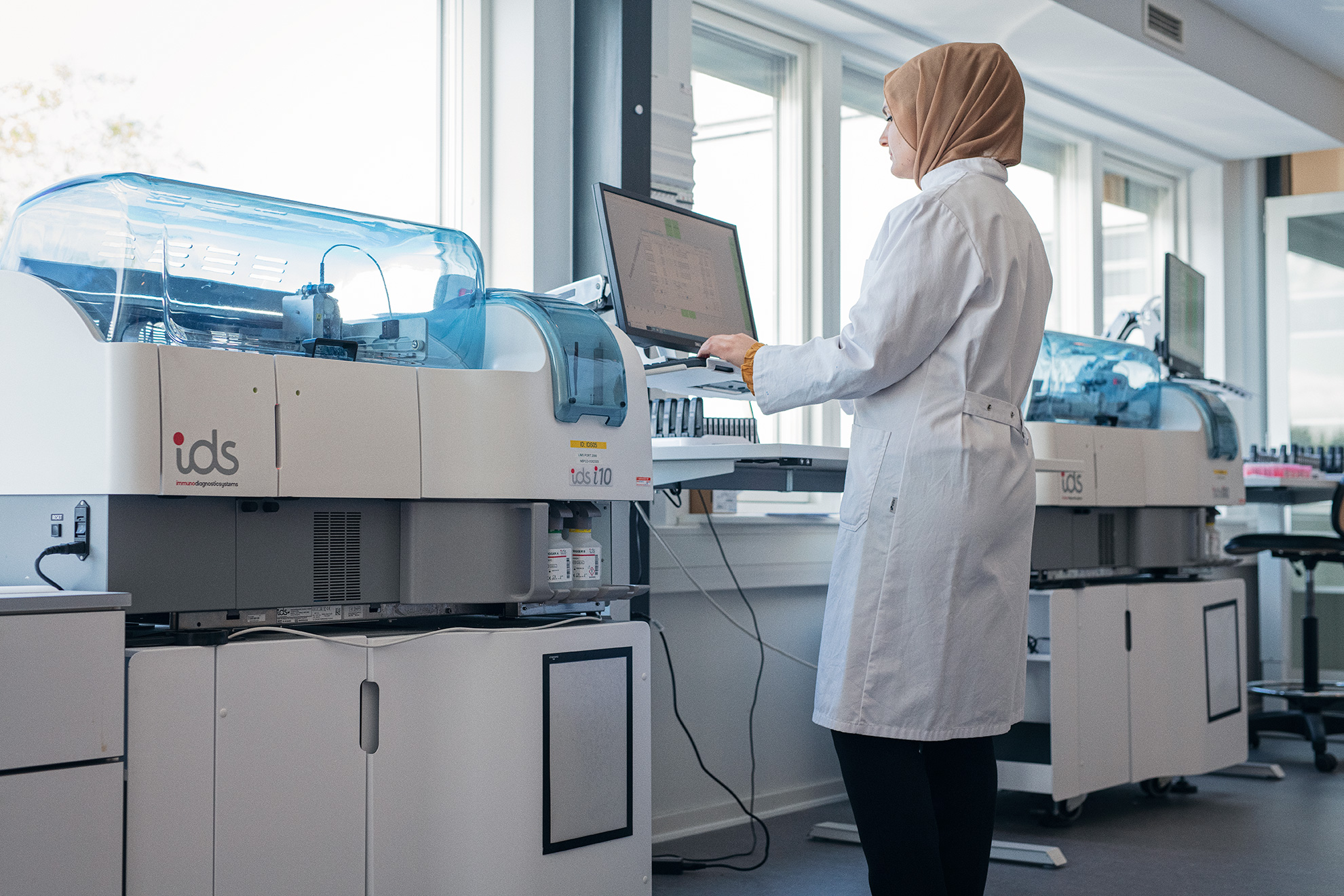
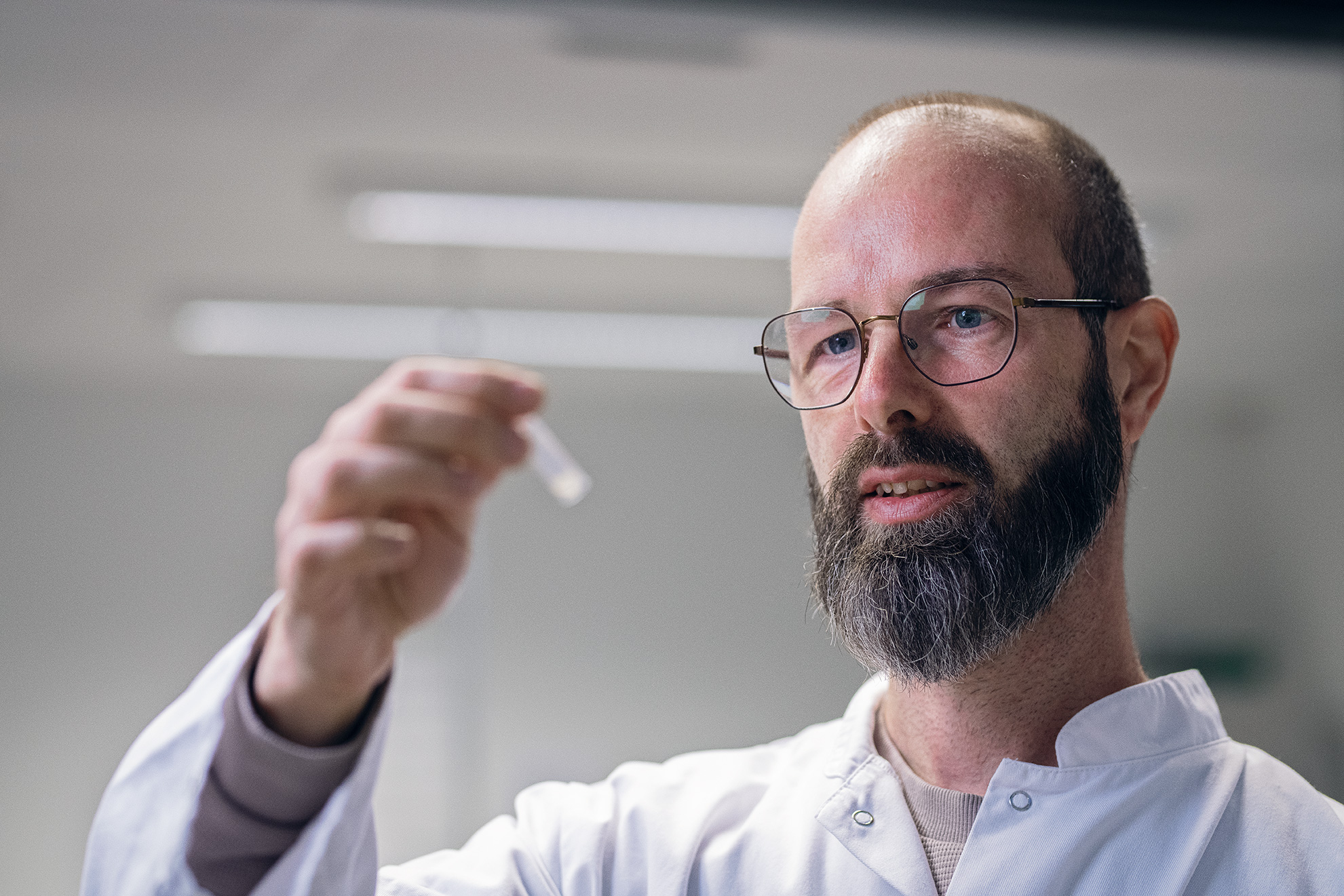
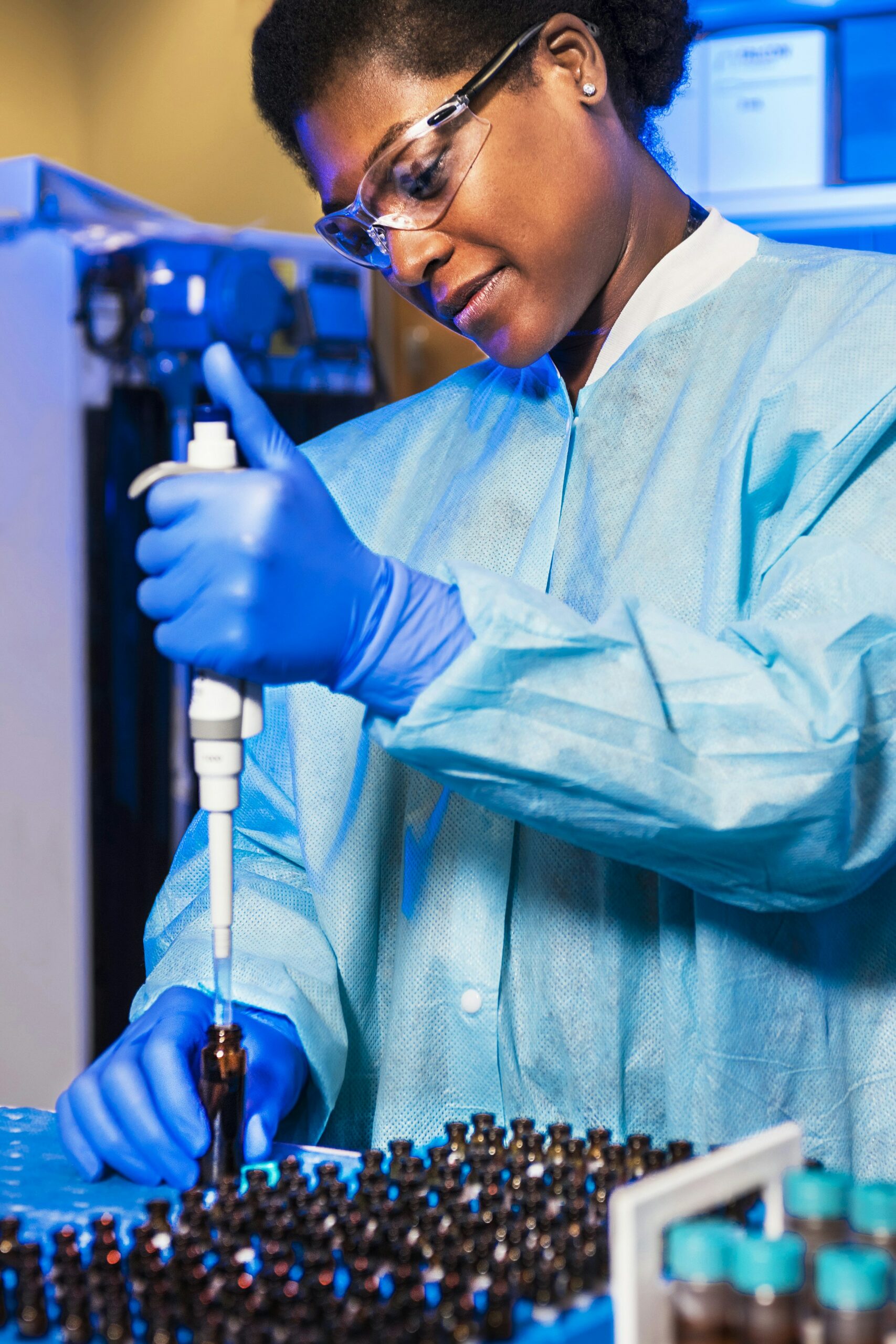
Nordic Bioscience’s ProteinFingerPrint Technology™ is backed by more than 30 years of research and used by world-leading pharmaceutical companies.
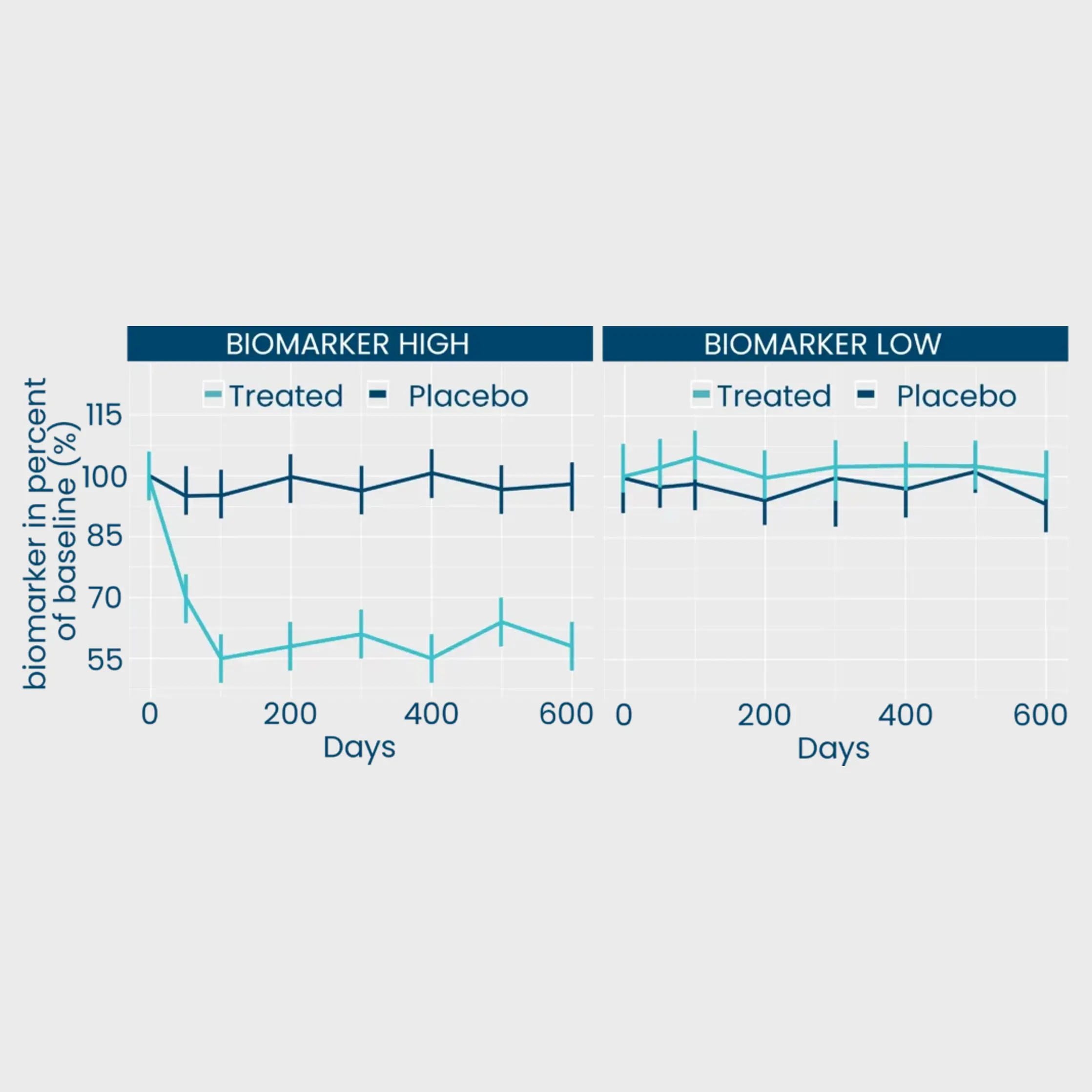
Our extracellular matrix-based approach quantifies disease activity with precision unmatched by omics providers, offering mechanistic and pharmacodynamic insights into tissue remodeling, a process crucial to understand when evaluating the effect of a therapeutic intervention.